gProfiler is a toolset for functional enrichment analysis and conversions of gene lists.
Biological data analysis often deals with lists of genes arising from various studies. The gProfiler toolset is widely used for finding biological categories enriched in gene lists, conversions between gene identifiers and mappings to their orthologs. The mission of gProfiler is to provide a reliable service based on up-to-date high quality data in a convenient manner across many evidence types, identifier spaces and organisms.
gProfiler relies on data retrieved from Ensembl database and fungi, plants or metazoa specific versions of Ensembl Genomes, and parasite specific data from WormBase ParaSite. In addition to Gene Ontology, supported data sources include KEGG, Reactome, WikiPathways, TRANSFAC, miRTarBase, Human Protein Atlas, CORUM and HP.
gProfiler supports close to 500 organisms and accepts hundreds of identifier types.
gProfiler has been supporting developers by offering a standardized access through public APIs. The usage of APIs was simplified by wrapping them into Python and R packages. The latter, gprofiler2 (which was published as a Conda Forge package named r-gprofiler2), was wrapped to be used as a set of Galaxy tools:
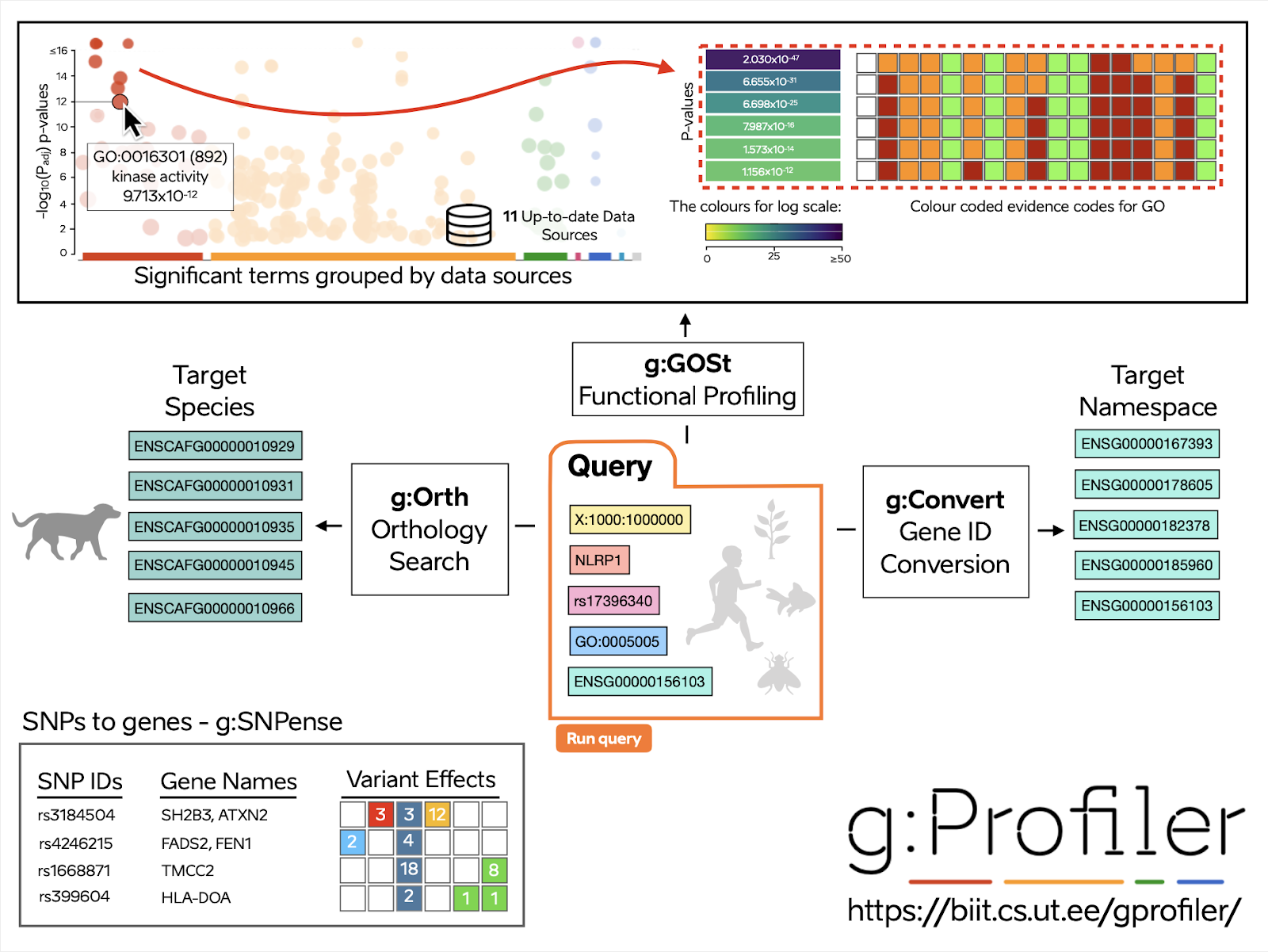
- gProfiler GOSt performs functional enrichment analysis, also known as over-representation analysis (ORA) or gene set enrichment analysis, on input gene list; in addition to the detailed tabular results, it can also create a Plotly graph with the Manhattan plot very similar to the one shown by the web tool.
- gProfiler Convert enables to convert between various gene, protein, microarray probe and numerous other types of namespaces
- gProfiler Orth translates gene identifiers between organisms
- gProfiler SNPense maps SNP rs-codes to gene names, chromosomal coordinates and variant effects
- gProfiler Random fetches a (not actually random) list of gene identifiers; useful for testing purposes.
gProfiler Galaxy toolset was admitted to the Galaxy Tools IUC. It is distributed via the ToolShed and can be installed as a bundle with the gProfiler tool suite repository.
gProfiler is part of the ELIXIR infrastructure. It belongs to the portfolio of ELIXIR Recommended Interoperability Resources.
gProfiler is developed and maintained in Estonia, at the University of Tartu, Institute of Computer Science, BIIT research group. It is an ELIXIR Estonia service.
